deep-purpose-tutorial
Deep-Purpose-Tutorial 
A low code tutorial for replicating DeepPurpose Library for Drug-Target Interaction (DTI) for the HackBio’2021 Internship as a part of Team Drug-Development-A

Drug-Dev-A will be using an open-source toolkit called DeepPurpose for Drug-Target Interaction (DTI) prediction which measures the binding affinity of drug molecules to the protein targets. What our team will be doing is replicating some of the DeepPurpose tutorialson our own google colaboratories for checking the reproducibility of the code samples and learning how this toolkit wraps deep learning (DL) models promising performances for DTI prediction to a comprehensive and easy-to-use DL library. Combining machine learned approaches to drug discovery is an effective way as shown by this web-app. This README.md contains all the necessary information to replicate the tutorials for DeepPurpose.
About DeepPurpose:
Working With DeepPurpose:
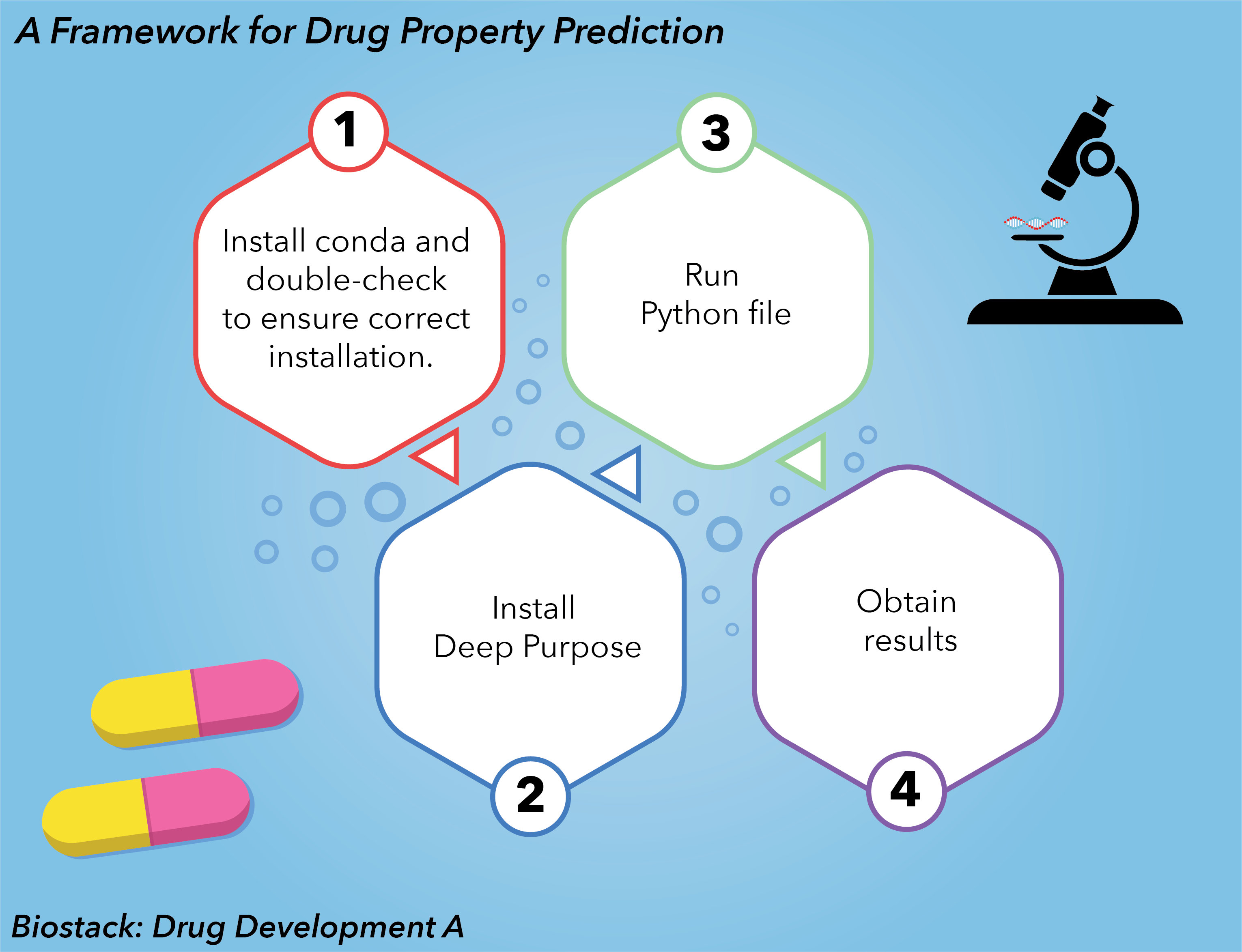
The work-horse for playing with DeepPurpose will be Google Colaboratory or Colab in short. We will be using Google Colaboratory to install DeepPurpose and it’s dependencies for running/eplicating the tutorials. Each of the steps should be run in different cells in Colab
- Google Colab does not come with Conda Environment installed. Conda Environment can be installed on google colab through ‘pip’:
!pip install -q condacolab import condacolab condacolab.install_anaconda() - Checking if Conda is working all fine:
import condacolab condacolab.check() !conda --version - All good! Now we will install DeepPurpose and it’s dependencies:
%%bash git clone https://github.com/kexinhuang12345/DeepPurpose.git ## Cloning the DeepPurpose Code Repository cd DeepPurpose ## Change Directory to DeepPurpose conda env create -f environment.yml # Creating New Environment source activate DeepPurpose - We have DeepPurpose, now installing some other dependencies:
conda install -c conda-forge rdkit conda install -c conda-forge notebook - Running a simple file for the tutorial, let’s call it drug.py. Contents of drug.py:
from DeepPurpose import oneliner #Importing the oneliner package from DeepPurpose.dataset import * #Importing the proprietary library of drugs oneliner.repurpose(*load_SARS_CoV2_Protease_3CL(), *load_antiviral_drugs(no_cid = True)) -
Run drug.py as
python drug.pyand voila you have retrieved a list of repurposing drugs from a proprietary library. - Output that you will get in a text file in your runtime:
----output---- Drug Repurposing Result for SARS-CoV2 3CL Protease +------+----------------------+------------------------+---------------+ | Rank | Drug Name | Target Name | Binding Score | +------+----------------------+------------------------+---------------+ | 1 | Sofosbuvir | SARS-CoV2 3CL Protease | 190.25 | | 2 | Daclatasvir | SARS-CoV2 3CL Protease | 214.58 | | 3 | Vicriviroc | SARS-CoV2 3CL Protease | 315.70 | | 4 | Simeprevir | SARS-CoV2 3CL Protease | 396.53 | | 5 | Etravirine | SARS-CoV2 3CL Protease | 409.34 | | 6 | Amantadine | SARS-CoV2 3CL Protease | 419.76 | | 7 | Letermovir | SARS-CoV2 3CL Protease | 460.28 | | 8 | Rilpivirine | SARS-CoV2 3CL Protease | 470.79 | | 9 | Darunavir | SARS-CoV2 3CL Protease | 472.24 | | 10 | Lopinavir | SARS-CoV2 3CL Protease | 473.01 | | 11 | Maraviroc | SARS-CoV2 3CL Protease | 474.86 | | 12 | Fosamprenavir | SARS-CoV2 3CL Protease | 487.45 | | 13 | Ritonavir | SARS-CoV2 3CL Protease | 492.19 | ....
Replicated DeepPurpose Tutorials:
Case Study 1(a): A Framework for Drug Target Interaction Prediction, with less than 10 lines of codes.
In addition to the DTI prediction, we also provide repurpose and virtual screening functions to rapidly generation predictions. The code for this replicated tutorial is available in a jupyter notebook interfaced with binder.
** Steps to walkthrough in this tutorial:**
- Import the DeepPurpose dependencies as we did earlier:
from DeepPurpose import DTI as models from DeepPurpose.utils import * from DeepPurpose.dataset import * - Loading the data:
# Load Data, an array of SMILES for drug, an array of Amino Acid Sequence for Target and an array of binding values/0-1 label. # e.g. ['Cc1ccc(CNS(=O)(=O)c2ccc(s2)S(N)(=O)=O)cc1', ...], ['MSHHWGYGKHNGPEHWHKDFPIAKGERQSPVDIDTH...', ...], [0.46, 0.49, ...] # In this example, BindingDB with Kd binding score is used. - Run the tutorial python file in the colab (instructions mentioned in the code comments): ``` X_drug, X_target, y = process_BindingDB(download_BindingDB(SAVE_PATH), y = ‘Kd’, binary = False, convert_to_log = True)
Type in the encoding names for drug/protein.
drug_encoding, target_encoding = ‘MPNN’, ‘Transformer’
Data processing, here we select cold protein split setup.
train, val, test = data_process(X_drug, X_target, y, drug_encoding, target_encoding, split_method=’cold_protein’, frac=[0.7,0.1,0.2])
Generate new model using default parameters; also allow model tuning via input parameters.
config = generate_config(drug_encoding, target_encoding, transformer_n_layer_target = 8) net = models.model_initialize(**config)
Train the new model.
Detailed output including a tidy table storing validation loss, metrics, AUC curves figures and etc. are stored in the ./result folder.
net.train(train, val, test)
or simply load pretrained model from a model directory path or reproduced model name such as DeepDTA
net = models.model_pretrained(MODEL_PATH_DIR or MODEL_NAME)
Repurpose using the trained model or pre-trained model
In this example, loading repurposing dataset using Broad Repurposing Hub and SARS-CoV 3CL Protease Target.
X_repurpose, drug_name, drug_cid = load_broad_repurposing_hub(SAVE_PATH) target, target_name = load_SARS_CoV_Protease_3CL()
_ = models.repurpose(X_repurpose, target, net, drug_name, target_name)
Virtual screening using the trained model or pre-trained model
X_repurpose, drug_name, target, target_name = [‘CCCCCCCOc1cccc(c1)C([O-])=O’, …], [‘16007391’, …], [‘MLARRKPVLPALTINPTIAEGPSPTSEGASEANLVDLQKKLEEL…’, …], [‘P36896’, ‘P00374’]
_ = models.virtual_screening(X_repurpose, target, net, drug_name, target_name)
**Figure: The Target SMILES used for Virtual Screening in the trained model**

**Link to the binder run for 1(a):** [](https://mybinder.org/v2/gh/ssiddhantsharma/deep-purpose-tutorial/HEAD?filepath=tutorial-notebooks%2FHackbio_Case_Study_1_(a)_A_Framework_for_Drug_Target_Interaction_Prediction.ipynb) <br>
**[Video-tutorial of 1(a) by a member of team through Loom-App](https://www.loom.com/share/1564269d811d410c9fcdcfdb2f55967a?sharedAppSource=personal_library)**
### Case Study 1(b): A Framework for Drug Property Prediction, with less than 10 lines of codes.
Many dataset is in the form of high throughput screening data, which have only drug and its activity score. It can be formulated as a drug property prediction task. We also provide a repurpose function to predict over large space of drugs. The code for this replicated tutorial is available in a jupyter notebook interfaced with binder.
**Link to the binder run for 1(b):** [](https://mybinder.org/v2/gh/ssiddhantsharma/deep-purpose-tutorial/HEAD?filepath=tutorial-notebooks%2FHackbio_Case_Study_1_(b)_A_Framework_for_Drug_Property_Prediction.ipynb) <br>
**[Video-tutorial of 1(b) by a member of team through Loom-App](https://www.loom.com/share/b38b55e16e184b45a3ae0fde3e3a9df0)**
### Case Study 2 (a): Antiviral Drugs Repurposing for SARS-CoV2 3CLPro, using One Line.
Given a new target sequence (e.g. SARS-CoV2 3CL Protease), retrieve a list of repurposing drugs from a curated drug library of 81 antiviral drugs. The Binding Score is the Kd values. The code for this replicated tutorial is available in a jupyter notebook interfaced with binder.
—-output—- Drug Repurposing Result for SARS-CoV2 3CL Protease +——+———————-+————————+—————+ | Rank | Drug Name | Target Name | Binding Score | +——+———————-+————————+—————+ | 1 | Sofosbuvir | SARS-CoV2 3CL Protease | 190.25 | | 2 | Daclatasvir | SARS-CoV2 3CL Protease | 214.58 | | 3 | Vicriviroc | SARS-CoV2 3CL Protease | 315.70 | | 4 | Simeprevir | SARS-CoV2 3CL Protease | 396.53 | | 5 | Etravirine | SARS-CoV2 3CL Protease | 409.34 | | 6 | Amantadine | SARS-CoV2 3CL Protease | 419.76 | | 7 | Letermovir | SARS-CoV2 3CL Protease | 460.28 | | 8 | Rilpivirine | SARS-CoV2 3CL Protease | 470.79 | | 9 | Darunavir | SARS-CoV2 3CL Protease | 472.24 | | 10 | Lopinavir | SARS-CoV2 3CL Protease | 473.01 | | 11 | Maraviroc | SARS-CoV2 3CL Protease | 474.86 | | 12 | Fosamprenavir | SARS-CoV2 3CL Protease | 487.45 | | 13 | Ritonavir | SARS-CoV2 3CL Protease | 492.19 | ….
**Link to the binder run for 2(a):** [](https://mybinder.org/v2/gh/ssiddhantsharma/deep-purpose-tutorial/HEAD?filepath=tutorial-notebooks%2FHackbio_Case_Study_2_(a)_Antiviral_Drugs_Repurposing_for_SARS_CoV2_3CLPro_using_One_Line.ipynb) <br>
**[Video-tutorial of 2(a) by a member of team through Loom-App](https://www.loom.com/share/7e3eac0a45144b9abb60bbea17383f27)**
### Case Study 2(b): Repurposing using Customized training data, with One Line.
Given a new target sequence (e.g. SARS-CoV 3CL Pro), training on new data (AID1706 Bioassay), and then retrieve a list of repurposing drugs from a proprietary library (e.g. antiviral drugs). The code for this replicated tutorial is available in a jupyter notebook interfaced with binder.
—-output—-
Drug Repurposing Result for SARS-CoV2 3CL Protease
+——+———————-+————————+—————+
| Rank | Drug Name | Target Name | Binding Score |
+——+———————-+————————+—————+
| 1 | Sofosbuvir | SARS-CoV2 3CL Protease | 190.25 |
| 2 | Daclatasvir | SARS-CoV2 3CL Protease | 214.58 |
| 3 | Vicriviroc | SARS-CoV2 3CL Protease | 315.70 |
| 4 | Simeprevir | SARS-CoV2 3CL Protease | 396.53 |
| 5 | Etravirine | SARS-CoV2 3CL Protease | 409.34 |
| 6 | Amantadine | SARS-CoV2 3CL Protease | 419.76 |
| 7 | Letermovir | SARS-CoV2 3CL Protease | 460.28 |
| 8 | Rilpivirine | SARS-CoV2 3CL Protease | 470.79 |
| 9 | Darunavir | SARS-CoV2 3CL Protease | 472.24 |
| 10 | Lopinavir | SARS-CoV2 3CL Protease | 473.01 |
| 11 | Maraviroc | SARS-CoV2 3CL Protease | 474.86 |
| 12 | Fosamprenavir | SARS-CoV2 3CL Protease | 487.45 |
| 13 | Ritonavir | SARS-CoV2 3CL Protease | 492.19 |
….
```
Link to the binder run for 2(b):
Video-tutorial of 2(b) by a member of team through Loom-App
DeepPurpose Reference:
If you found this package interactive, please see their publication DeepPurpose:

Contributors (Slack Usernames)
All people contributed to running their copy of DeepPurpose on Colab atleast once.
Case Study 1(a):
- @Emon : Installed resources locally and on google colab, replicated Case Study 1(a).
- @Hameed : Installed resources locally and on google colab, replicated Case Study 1(a) and recording the tutorial video.
- @Sanjana : Installed resources locally and on google colab, replicated Case Study 1(a) and confirmation of final reproducibility.
Case Study 1(b):
- @eshaan : Installed resources locally and on google colab, replicated Case Study 1(b) and recording the tutorial video.
- @Ayush : Installed resources locally and on google colab, replicated Case Study 1(b) and troubleshooting of the whole team’s errors.
- @Anjali : Installed resources locally and on google colab, replicated Case Study 1(b) and confirmation of final reproducibility and designed the DeepPurpose Banner.
Case Study 2(a):
- @UmasriSankarlal : Installed resources locally and on google colab, replicated Case Study 2(a).
- @jovel : Installed resources locally and on google colab, replicated Case Study 2(a) and recording the tutorial video.
- @Ayesha : Installed resources locally and on google colab, replicated Case Study 2(a) and confirmation of final reproducibility.
Case Study 2(b):
- @Benson: Installed resources locally and on google colab, replicated Case Study 2(b) and recording the tutorial video.
- @Dolu : Installed resources locally and on google colab, replicated Case Study 2(b) and troubleshooting of the team’s errors.
- @Maurya : Installed resources locally and on google colab, replicated Case Study 2(b) and confirmation of final reproducibility and DeepPurpose Banner.
Contributors (Slack Usernames)
- @Jass: All graphical figures in Adobe Suite and setting up Binder wrapping for the jupyter notebooks of the tutorials.
- @Siddhant: Idea Generation and Github Repository Management.
🎨 Future Improvements/Ideas:
One can try to benchmark DeepPurpose with the very new (August 2021) library, TorchDrug, might give something good to compare to. More about installing TorchDrug here. In case of any questions, please send to siddhaantsharma.ss@gmail.com




